
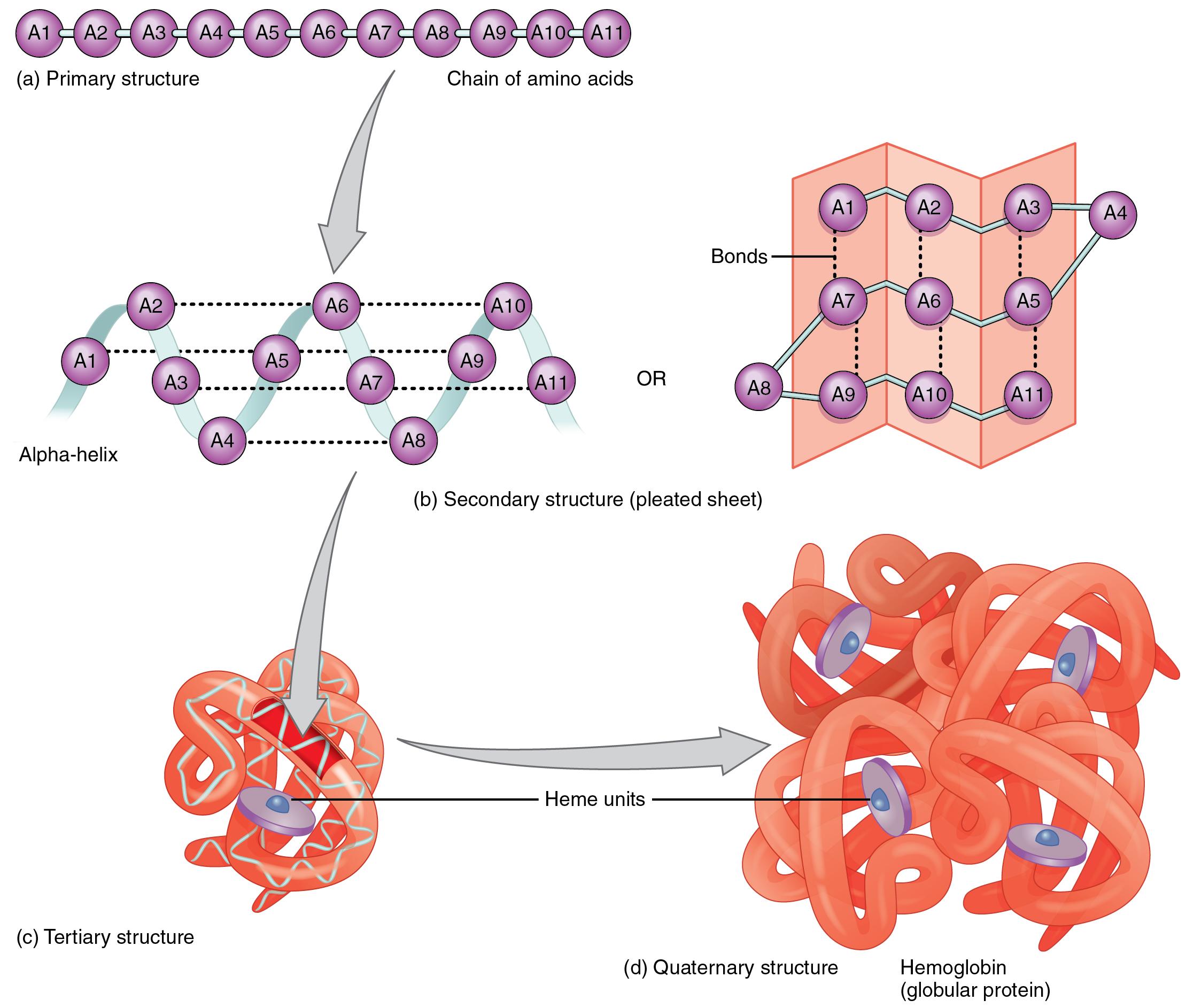
This tutorial review describes the background to the technique of CD spectroscopy and good practice methods for high quality data collection. More than 24 000 papers published in the past decade have included CD characterisations of proteins many of those studies have also included other complementary chemical, biophysical, and computational chemistry methods. RF Diffusion uses this strategy to turn clouds of disconnected atoms into coherent protein structures.Circular dichroism (CD) spectroscopy is a widely-used method in biochemistry, structural biology and pharmaceutical chemistry. Additional software guides this de-noising process. Diffusion models begin with grainy bits of static and gradually remove noise until a clear picture is formed. Image generation tools like DALL-E produces high-quality images that have never existed before using a diffusion model, which is a machine-learning algorithm that specializes in adding and removing noise. Using RF Diffusion, we find that as little as one design must be tested. With prior design methods, tens of thousands of protein designs may have to be tested before finding a single one that performs as intended. RoseTTAFold Diffusion (RF Diffusion) is a guided diffusion model for generating new proteins. It outperforms existing protein design methods across a broad range of problems and has been used to generate ultra-high affinity binders through pure computation and a series of novel symmetric assemblies that we have experimentally validated via electron microscopy. And we have shown that RoseTTAFold can be used to build models of complex biological assemblies in a fraction of the time previously required. We also generated structures directly relevant to human health, including for proteins associated with problematic lipid metabolism, inflammation disorders, and cancer cell growth. In this architecture, one-, two-, and three-dimensional information flows back and forth, allowing the network to collectively reason about the relationship between a protein’s chemical parts and its folded structure.Īs reported in Science, our team has used RoseTTAFold to compute hundreds of new protein structures, including many poorly understood proteins from the human genome.
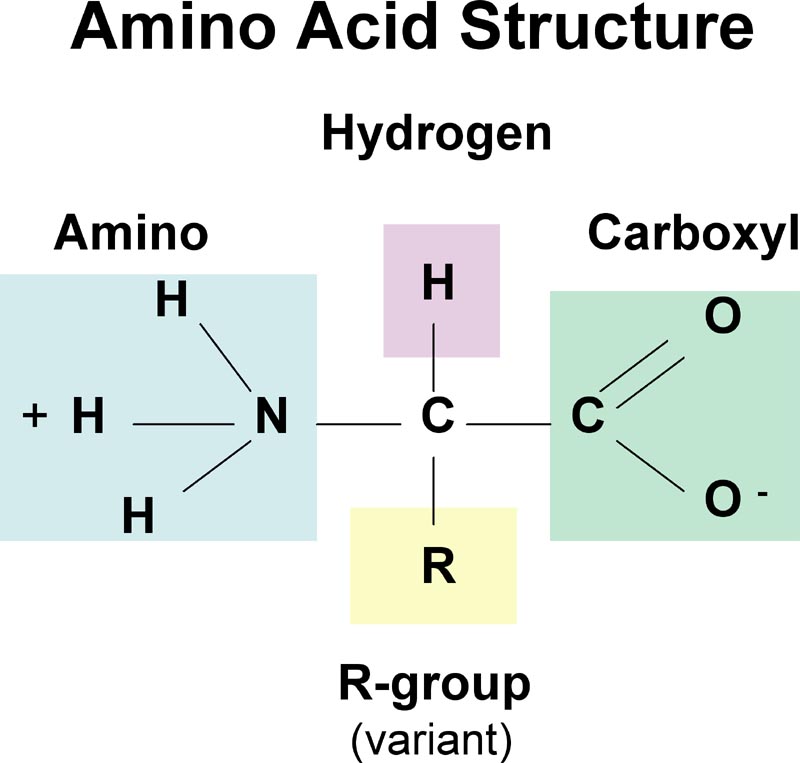
RoseTTAFold is a three-track neural network, meaning it simultaneously considers patterns in protein sequences, how a protein’s amino acids interact with one another, and a protein’s possible three-dimensional structure.
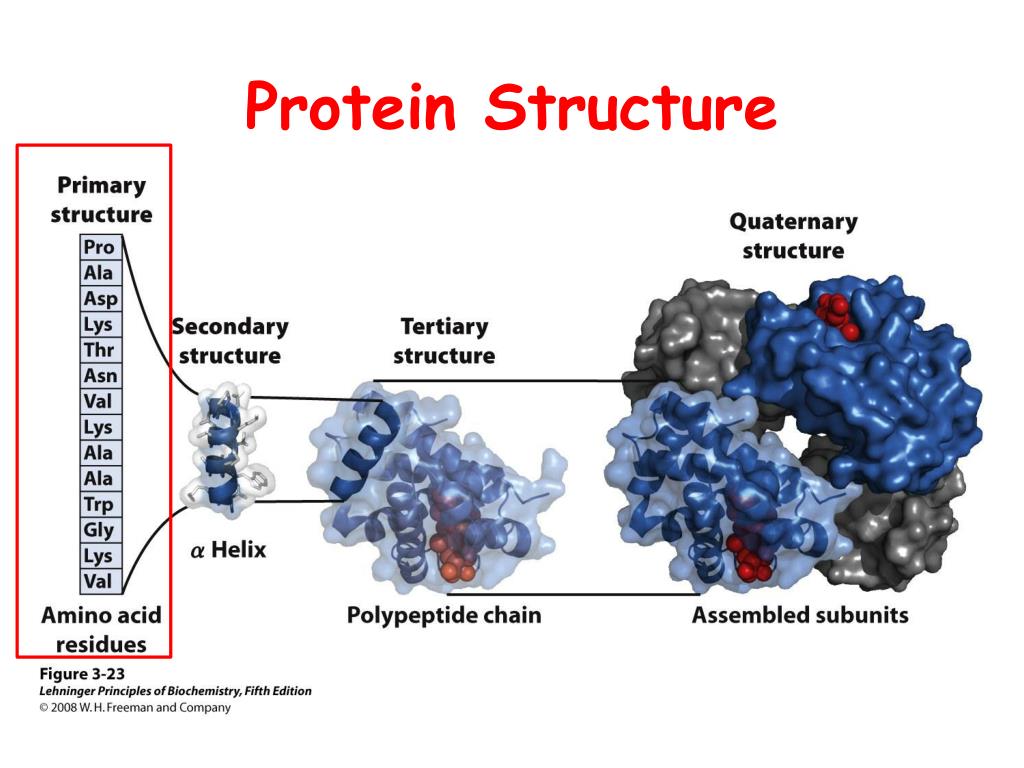
With RoseTTAFold, a protein structure can be computed in as little as ten minutes on a single gaming computer. Without the aid of such software, it can take years of laboratory work to determine the structure of just one protein. RoseTTAFold is a software tool that uses deep learning to quickly and accurately predict protein structures based on amino acid sequences alone.


 0 kommentar(er)
0 kommentar(er)
